- microfilaments (composed of an actin polymer) and
- microtubules (composed of a polymer of tubulin.
Actin
The monomeric unit of actin is called G-actin (globular actin) and the polymer is known as F-actin (filamentous actin). Filaments of Factin comprise the smallest filaments of cells known as microfilaments (Figure 2.101). Actin is essential for muscular contraction and also has diverse roles in cellular signaling and maintenance of cell junctions. In conjunction with other proteins, actin has numerous interactions with the cell membrane. The β- and γ-forms of actin are components of the cytoskeleton and facilitate motility inside of cells. α-actin is important in muscle tissues, where it is used by myosin in the mechanical process of contraction (See HERE).
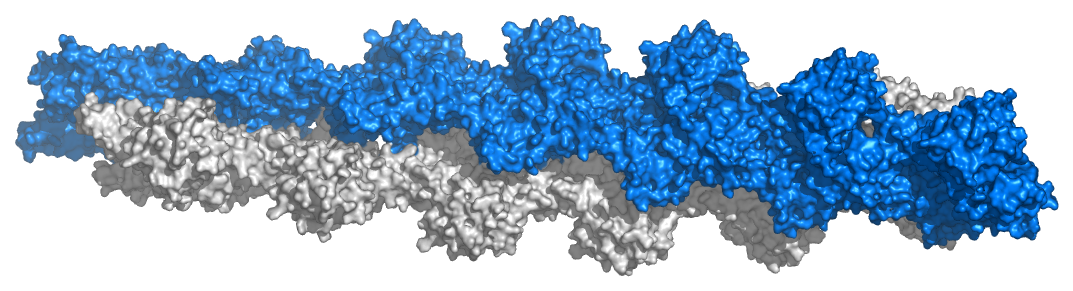
Figure 2.101 – Model of actin filaments. Image used with permission (CC BY-SA 3.0; Thomas Splettstoesser).
Monomeric and polymeric forms of actin play roles in cellular activities relating to motion. Two parallel F-actin strands can pair with each other and create a double helical structure with 2.17 subunits per turn of the helix. Helical F-actin in muscles contains tropomyosin, which covers the actin binding sites for myosin in resting muscles to prevent contraction. Other proteins bound to actin muscle filaments include the troponins (I, T, and C).
Actin Cellular Action
Examples of actin action at the cellular level include cell motility, cytokinesis, intracellular transport of vesicles and organelles, and cell shape. Each actin monomer is bound to a molecule of ATP or ADP and the presence of one of these is essential for proper G-actin functioning.
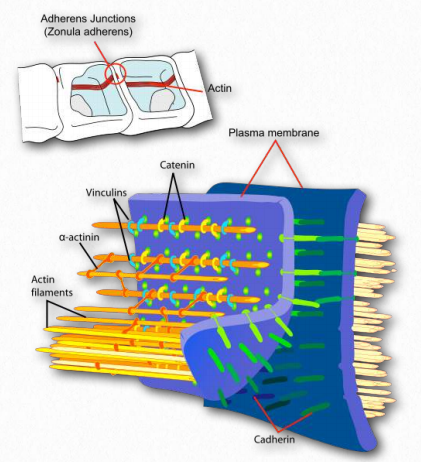
Figure 2.102 – Attachment of actin at the cell membrane complex known as the adherens junction Wikipedia
The role of ATP
In the monomer, actin is more commonly bound to ATP, whereas in the filaments, it is typically bound to ADP. Actin is an inefficient ATPase, breaking the molecule down slowly, but the catalysis speeds up as much as 40,000 fold when the monomer begins to polymerize. Actin also has a binding site for divalent cations – either calcium or magnesium. F-
Actin binds to structural proteins at the adherens junction (Figure 2.102). These include α-actinin, vinculin (provides a membrane connection and connections to the catenins and cadherin).
Polymerization
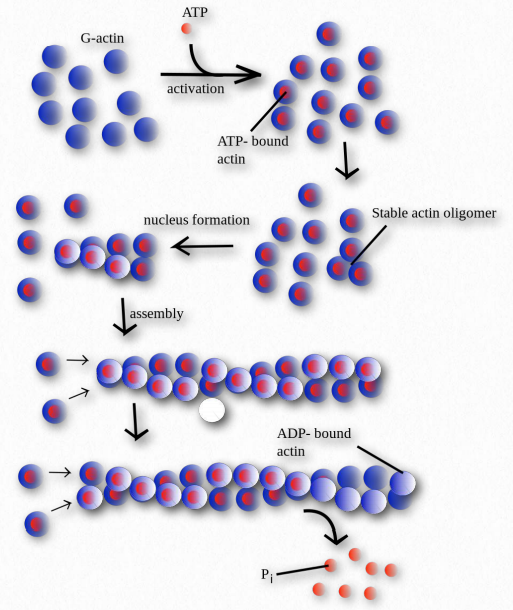
Figure 2.103 – Polymerization of G-actin monomers into Factin polymers
Polymerization of actin begins with a nucleating event (Figure 2.103). One factor known to affect the process is known as the Arp 2/3 complex. It does this by mimicking an actin dimer, starting an autocatalytic process of actin assembly. The Arp 2/3 complex plays roles both in the initiation of polymerization of new actin filaments as well as the formation of branches in the filaments.
Two proteins play roles in modulating polymer growth. Thymosin functions on the end of actin filaments to control growth. Profilin works on G-actin monomers exchanging ADP for ATP, promoting addition of monomers to a growing chain.
F-actin filaments are held together by relatively weak bonds compared to the covalent bonds of the monomers of nucleic acids, thus allowing for easier disassembly when desired. Actin’s amino acid sequence is optimized, having diverged only a relatively small amount (20%) between algae and humans. Mutations in the actin gene result in muscular diseases and/or deafness.
Tubulin
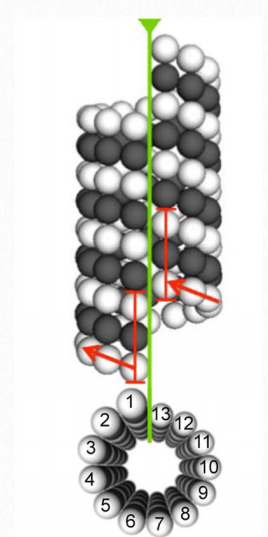
Figure 2.104 Microtubule structure Wikipedia
Tubulin proteins are the monomeric building blocks of eukaryotic microtubules (Figure 2.104 & 2.105). Bacterial (TubZ) and archaeon (FtsZ) equivalents are known. The α-tubulin and β-tubulin proteins polymerize to make microtubule structures in the cytoplasm of cells. Microtubules are major components of the cytoskeleton of eukaryotic cells, providing structural support, transport within the cell, and functions necessary for segregation of DNAs during cell division.
Dimerization of the α-tubulin and β-tubulin proteins is necessary for polymerization and requires that the subunits bind to GTP. Microtubules only grow in one direction. β- tubulin is found on the plus end of the tubule (growth end = plus end) and α-tubulin is exposed on the other end (non-growth end = minus end). Dimers of α-tubulin/β-tubulin are incorporated into growing microtubules in this orientation. If a dimer is bound to GDP instead of GTP, it tends to be unstable and fall apart, whereas those bound to GTP stably assemble into microtubules.
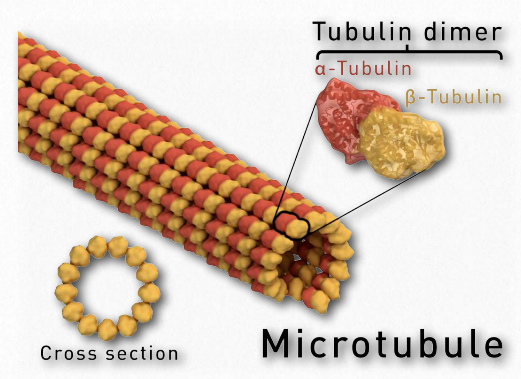
Figure 2.105 – Microtubule anatomy Wikipedia
Microtubules
Microtubules, along with microfilaments and intermediate filaments (see HERE) constitute the cytoskeleton of cells. Found in the cytoplasm, they are found in eukaryotic cells, as well as some bacteria. Microtubules help to give cells structure. They comprise the inner structure of flagella and cilia and provide rail-like surfaces for the transport of materials within cells.
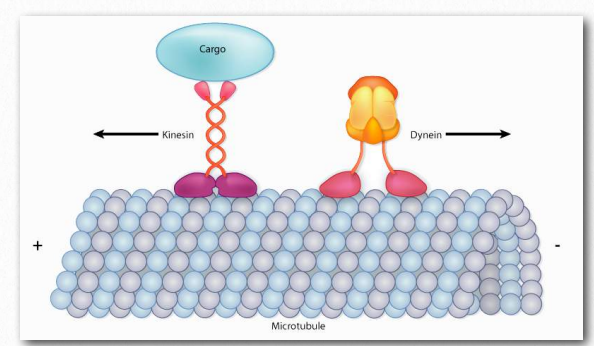
Figure 2.106 – Kinesin and dynein “walk” along microtubules, but move in opposite directions Image by Aleia Kim
Polymerization of α- tubulin and β-tubulin to form microtubules occurs after a nucleating event. Individual units get arranged in microtubule organizing centers (MTOCs), an example of which is the centrosome. Centrosomes are focal points of connection of microtubules. Basal bodies of cilia and flagella also help to organize microtubules.
Motor proteins
From the transport of materials within a cell to the process of cytokinesis where one cell splits into two in mitosis, a cell has needs for motion at the molecular level. Secretory vesicles and organelles must be transported. Chromosomes must be separated in mitosis and meiosis.
The proteins dynein and kinesin (Figure 2.106) are necessary for intracellular movement. These motor proteins facilitate the movement of materials inside of cells along microtubule “rails”. These motor proteins are able to move along a portion of the cytoskeleton by converting chemical energy into motion with the hydrolysis of ATP. An exception is flagellar rotation, which uses energy provided from a gradient created by a proton pump.
Kinesins and dyneins
As noted, kinesins and dyneins navigate in cells on microtubule tracks (Figure 2.108 & Movie 2.4). Most kinesins move in the direction of the synthesis of the microtubule (+ end movement), which is generally away from the cell center and the opposite direction of movement of dyneins, which are said to do retrograde transport toward the cell center. Both proteins provide movement functions necessary for the processes of mitosis and meiosis. These include spindle formation, chromosome separation, and shuttling of organelles, such as the mitochondria, Golgi apparatuses, and vesicles.
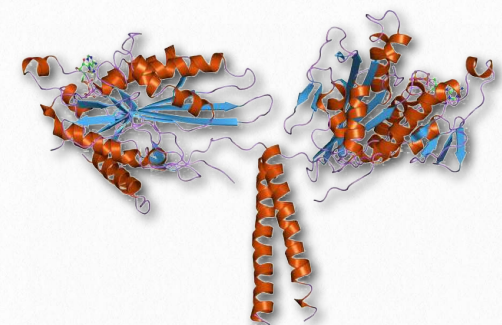
Figure 2.107 – Kinesin. “Feet” are at the top.
Kinesins are comprised of two heavy chains and two light chains. The head motor domains of heavy chains (in the feet) use energy of ATP hydrolysis to do mechanical work for the movement along the microtubules. There are at least fourteen distinct kinesin families and probably many related ones in addition.
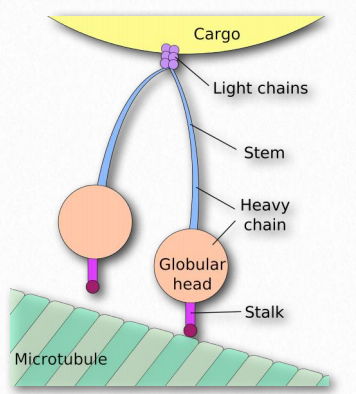
Figure 2.108 – Nomenclature of dynein. The “feet” of Figure 2.105 are the stalk and globular head of the structure here. Wikipedia
Dyneins are placed into two groups – cytoplasmic and axonemal (also called ciliary or flagellar dyneins – Figure 2.109). Dyneins are more complex in structure than kinesins with many small polypeptide units. Notably, plants do not have dynein motor proteins, but do contain kinesins.
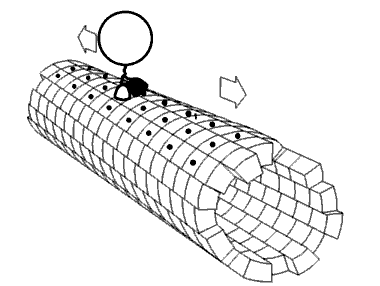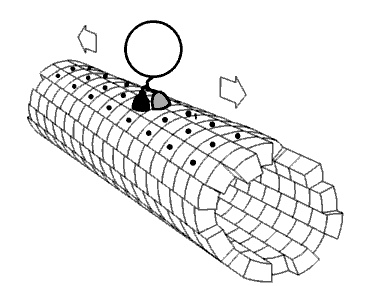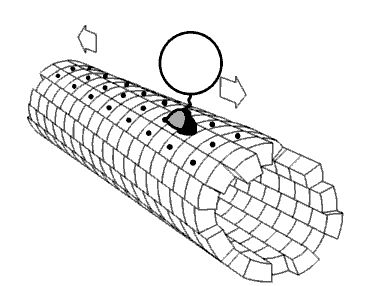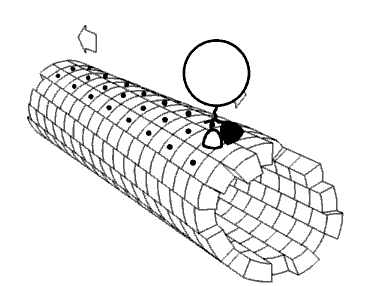
Movie 2.4 The motor protein kinesin walking down a microtubule. Image used with permission (Public Domain; zp706).
Myosin
An important group of motor proteins in the cell is the myosins. Like kinesins and dyneins, myosins use energy from hydrolysis of ATP for movement. In this case, the movement is mostly not along microtubules, but rather along microfilaments comprised of a polymer of actin (F-actin). Movement of myosin on actin is best known as the driving force for muscular contraction. Myosins are a huge family of proteins, all of which bind to actin and all of which involve motion. Eighteen different classes of myosin proteins are known.
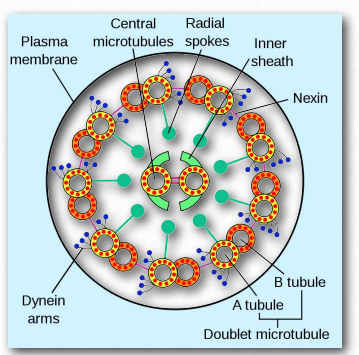
Figure 2.109 – Dynein in an axoneme Wikipedia
Myosin II is the form responsible for generating muscle contraction. It is an elongated protein formed from two heavy chains with motor heads and two light chains. Each myosin motor head binds actin and has an ATP binding site. The myosin heads bind and hydrolyze ATP. This hydrolysis produces the energy necessary for myosin to walk toward the plus end of an actin filament.
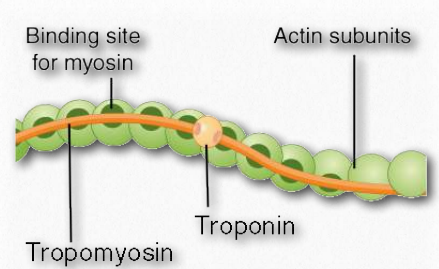
Figure 2.110 – Actin filament anatomy Wikipedia
Non-muscle myosin IIs provide contraction needed to power the action of cytokinesis. Other myosin proteins are involved in movement of non-muscle cells. Myosin I is involved in intracellular organization. Myosin V performs vesicle and organelle transport. Myosin XI provides movement along cellular microfilament networks to facilitate organelle and cytoplasmic streaming in a particular direction.
Structure
Myosins have six subunits, two heavy chains and four light chains. Myosin proteins have domains frequently described as a head and a tail (Figure 2.111). Some also describe an intermediate hinge region as a neck. The head portion of myosin is the part that binds to actin. It uses energy from ATP hydrolysis to move along the actin filaments. In muscles, myosin proteins form aggregated structures referred to as thick filaments. Movements are directional.
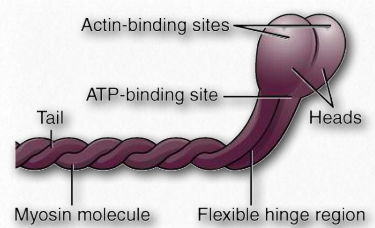
Figure 2.111 – Myosin protein anatomy Wikipedia
Structural considerations of muscular contraction
Before we discuss the steps in the process of muscular contraction, it is important to describe anatomical aspects of muscles and nomenclature.
There are three types of muscle tissue – skeletal (striated), smooth, and (in vertebrates) cardiac. We shall concern ourselves mostly here with skeletal muscle tissue. Muscles may be activated by the central nervous system or, in the case of smooth and cardiac muscles, may contract involuntarily. Skeletal muscles may be slow twitch or fast twitch.
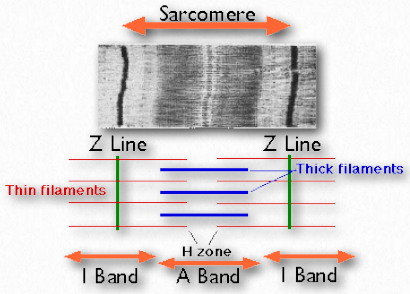
Figure 2.112 – Structural components of muscle. Wikipedia
Sarcomeres
Sarcomeres are described as the basic units comprising striated muscles and are comprised of thick (myosin) and thin (actin) filaments and a protein called titin. The filaments slide past each other in muscular contraction and then backwards in muscular relaxation. They are not found in smooth muscles.
Under the microscope, a sarcomere is the region between two Z-lines of striated muscle tissue (Figure 2.112). The Z-line is the distinct, narrow, dark region in the middle of an I-band. Within the sarcomere is an entire Aband with its central H-zone. Within the Hzone are located tails of myosin fibers, with the head pointed outwards from there projecting all the way to the I-band. The outside of the Aband is the darkest and it gets lighter moving towards the center.
Within the Iband are located thin filaments that are not occupied with thick myosin filaments. The Aband contains intact thick filaments overlaying thin filaments except in the central H zone, which contains only thick filaments. In the center of the H-zone is a line, known as the M-line. It contains connecting elements of the cellular cytoskeleton. In muscular contraction, myosin heads walk along pulling their tails over the actin thin filaments, using energy from hydrolysis of ATP and pulling them towards the center of the sarcomere.
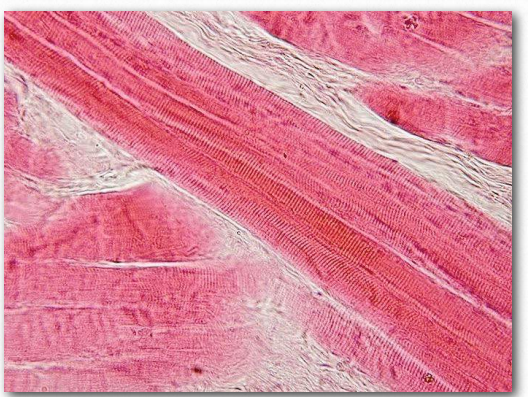
Figure 2.113 – Skeletal muscle longitudinal section. Wikipedia
Sarcolemma
The sarcolemma (also known as the myolemma) is to muscle cells what the plasma membrane is to other eukaryotic cells – a barrier between inside and outside. It contains a lipid bilayer and a glycocalyx on the outside of it. The glyocalyx contains polysaccharides and connects with the basement membrane. The basement membrane serves a s scaffolding to connect muscle fibers to. This connection is made by transmembrane proteins bridging the actin cytoskeleton on the inside of the cell with the basement membrane on the outside. On the ends of the muscle fibers, each sarcolemma fuses with a tendon fiber and these, in turn, adhere to bones.
Sarcoplasmic reticulum
The sarcoplasmic reticulum (Figure 2.114) is a name for the structure found within muscle cells that is similar to the smooth endoplasmic reticulum found in other cells. It contains a specialized set of proteins to meet needs unique to muscle cells. The organelle largely serves as a calcium “battery,” releasing stored calcium to initiate muscular contraction when stimulated and taking up calcium when signaled at the end of the contraction cycle. It accomplishes these tasks using calcium ion channels for release of the ion and specific calcium ion pumps to take it up.
Movement direction
All myosins but myosin VI move towards the + end (the growing end) of the microfilament. The neck portion serves to link the head and the tail. It also a binding site for myosin light chain proteins that form part of a macromolecular complex with regulatory functions. The tail is the point of attachment of molecules or other “cargo” being moved. It can also connect with other myosin subunits and may have a role to play in controlling movement.
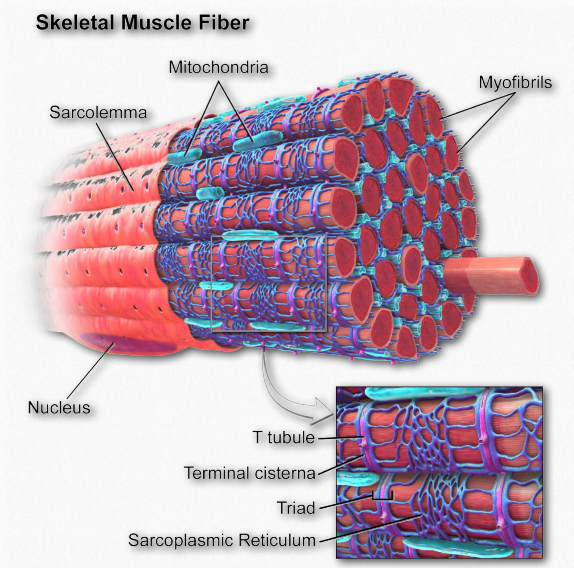
Figure 2.114 – Anatomy of a muscle fiber Wikipedia
Muscular contraction
The sliding filament model has been proposed to describe the process of muscular tension/contraction. In this process a repeating set of actions slide a thin actin filament over a thick myosin filament as a means of creating tension/ shortening of the muscle fiber.
Steps in the process occur as follows:
A. A signal from the central nervous system (action potential) arrives at a motor neuron, which it transmits towards the neuromuscular junction (see more on the neurotransmission part of the process HERE)
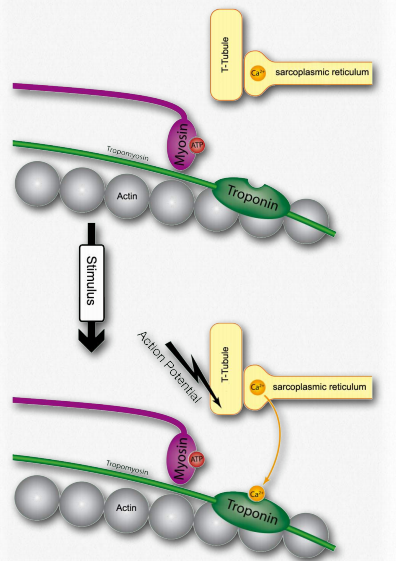
Figure 2.115 – 1. Activation of a muscle cell by release of calcium (step H) Wikipedia
B. At the end of the axon, the nerve signal stimulates the opening of calcium channels at the axon terminus causing calcium to flow into the terminal.
C. Movement of calcium into the axon of the nerve causes acetylcholine (a neurotransmitter) in synaptic vesicles to fuse with the plasma membrane. This causes the acetylcholine to be expelled into the synaptic cleft between the axon and the adjacent skeletal muscle fiber.
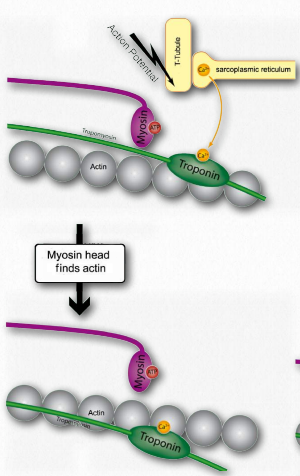
Figure 2.116 – 2. Calcium binding by troponin allows myosin to access actin sites (I). Wikipedia
D. Acetylcholine diffuses across the synapse and then binds to nicotinic acetylcholine receptors on the neuromuscular junction, activating them.
E. Activation of the receptor stimulates opening gates of sodium and potassium channels, allowing sodium to move into the cell and potassium to exit. The polarity of the membrane of the muscle cell (called a sarcolemma – Figure 2.111) changes rapidly (called the end plate potential).
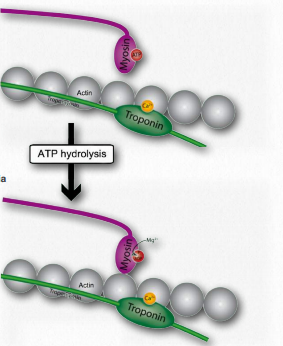
Figure 2.117 – 3. ATP cleavage by myosin allows actin attachment (J) Wikipedia
F. Change in the end plate potential results in opening of voltage sensitive ion channels specific for sodium or potassium only to Figure 2.117 – 3. ATP cleavage by myosin allows actin attachment (J) Wikipediaopen, creating an action potential (voltage change) that spreads throughout the cell in all directions.
G. The spreading action potential depolarizes the inner muscle fiber and opens calcium channels on the sarcoplasmic reticulum (Figure 2.115).
H. Calcium released from the sarcoplasmic reticulum binds to troponin on the actin filaments (Figure 2.115).
I. Troponin alters the structure of the tropomyosin to which is it bound. This causes tropomyosin to move slightly, allowing access to myosin binding sites on the microfilament (also called thin filament) that it was covering (Figure 2.116).
J. Myosin (bound to ATP) cleaves the ATP to ADP and Pi, which it holds onto in its head region and then attaches itself to the exposed binding sites on the thin filaments causing inorganic phosphate to be released from the myosin followed by ADP (Figure 2.117).
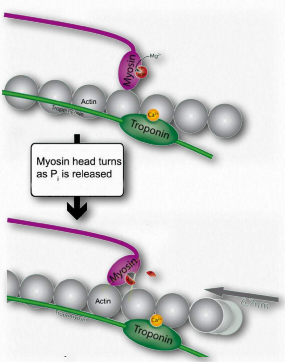
Figure 2.118 – 4. Release of Pi causes myosin hinge to bend. Thin filament pulled left (K). Wikipedia
K. Release of ADP and Pi is tightly coupled to a bending of the myosin hinge, resulting in what is called the power stroke. This causes the thin filament to move relative to the thick fibers of myosin (Figures 2.118 & 2.119).
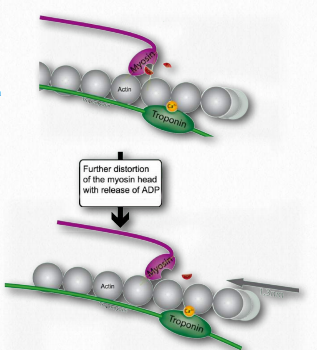
Figure 2.119 – 5. Release of ADP favors further bending of hinge and movement of thin filament leftward (K). Wikipedia
L. Such movement of the thin filaments causes the Z lines to be pulled closer to each other. This results in shortening of the sarcomere as a whole (Figure 2.122) and narrowing of the I band and the H zones (Figure 2.123). M. If ATP is available, it binds to myosin, allowing it to let go of the actin (Figures 2.120 & 2.121). If ATP is not available, the muscle will remain locked in this state. This is the cause of rigor mortis in death – contraction without release of muscles
.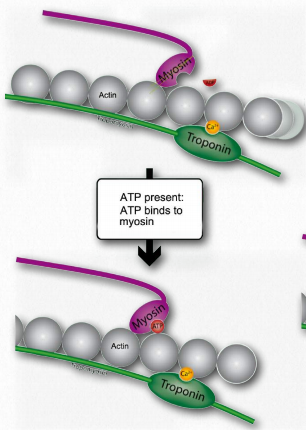
Figure 2.120 – When ATP is present, it binds to myosin (M). Wikipedia
N. After myosin has bound the ATP, it hydrolyzes it, producing ADP and Pi, which are held by the head. Hydrolysis of ATP resets the hinge region to its original state, unbending it. This unbent state is also referred to as the cocked position.
O.If tropomyosin is still permitting access to binding sites on actin, the process repeats so long as ATP is available and calcium remains at a high enough concentration to permit it to bond to troponin.
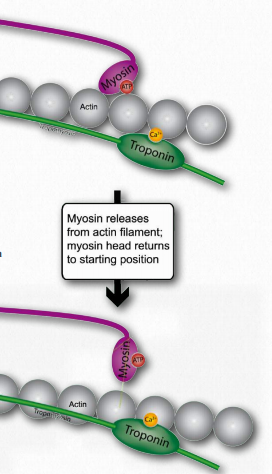
Figure 2.121 – Binding of ATP favors release of myosin from actin site (N) Wikipedia
Relaxation of the muscle tension occurs as the action potential in the muscle cell dissipates. This happens because all of the following things happen 1) the nerve signal stops; 2) the neurotransmitter is degraded by the enzyme acetylcholinesterase; and 3) the calcium concentration declines because it is taken up by the sarcoplasmic reticulum.
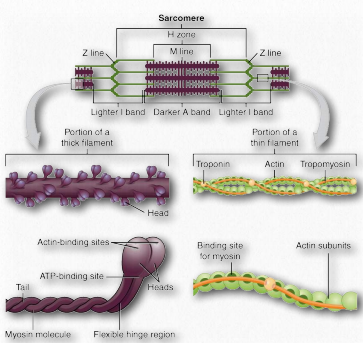
Figure 2.122 – Sarcomere Anatomy Wikipedia
It should be noted that the sarcoplasmic reticulum is always taking up calcium. Only when its calcium gates are opened by the action potential is it unable to reduce cellular calcium concentration. As the action potential decreases, then the calcium gates close and the sarcoplasmic reticulum “catches up” and cellular calcium concentrations fall. At that point troponin releases calcium, tropomyosin goes back to covering myosin binding sites on actin, myosin loses its attachment to actin and the thin filaments slide back to their original positions relative to the myosin thick filaments.
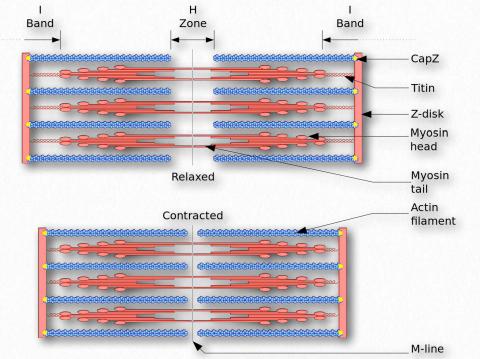
Figure 2.123 – The Sliding Filament Model of Muscular Contraction Wikipedia
Tropomyosin
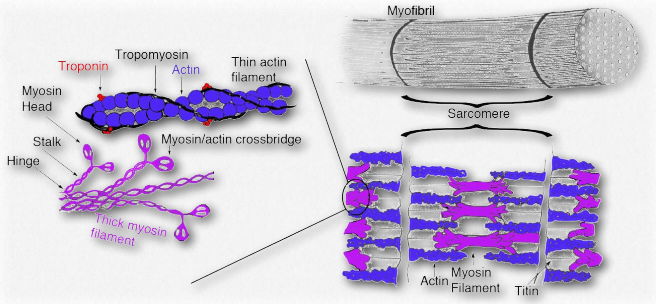
Figure 2.124 – Tropomyosin and troponin in muscle anatomy Wikipedia
Tropomyosins are proteins that interact with actin thin filaments to help regulate their roles in movement, both in muscle cells and non-muscle cells (Figure 2.124). Tropomyosins interact to form head-to-toe dimers and perch along the α-helical groove of an actin filament. The isoforms of tropomyosin that are in muscle cells control interactions between myosin and the actin filament within the sarcomere and help to regulate contraction of the muscle. In other cells, nonmuscle tropomyosins help to regulate the cytoskeleton’s functions.
The interactions of tropomyosin with the cytoskeleton are considerably more complicated than what occurs in muscle cells. Muscle cells have five tropomyosin isoforms, but in the cytoskeleton of non-muscle cells, there are over 40 tropomyosins.
Troponin
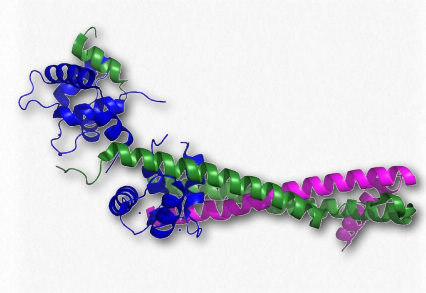
Figure 2.125 – Troponin complex of muscle. Blue = troponin C, magenta = troponin T , green = troponin I
The troponins involved in muscular contraction are actually a complex of three proteins known as troponin I, troponin C, and troponin T (Figure 2.125). They associate with each other and with tropomyosin on actin filaments to help regulate the process of muscular contraction. Troponin I prevents binding of myosin’s head to actin and thus prevents the most important step in contraction.
Troponin C is a unit that binds to calcium ions. Troponin T is responsible for binding all three proteins to tropomyosin. Troponins in the bloodstream are indicative of heart disorders. Elevation of troponins in the blood occurs after a myocardial infarction and can remain high for up to two weeks.
Actinin
Actinin is a skeletal muscle protein that attaches filaments of actin to Z-lines of skeletal muscle cells. In smooth muscle cells, it also connects actin to dense bodies.
Titin
Titin (also known as connectin) is the molecular equivalent of a spring that provides striated muscle cells with elasticity. It is the third most abundant protein in muscle cells. The protein is enormous, with 244 folded individual protein domains spread across 363 exons (largest known number), with the largest known exon (17,106 base pairs long), and it is the largest protein known (27,000 to 33,000 amino acids, depending on splicing).
Unstructured sequences
The folded protein domains are linked together by unstructured sequences. The unstructured regions of the protein allow for unfolding when stretching occurs and refolding upon relaxation. Titin connects the M and Z lines in the sarcomere (Figure 2.123). Tension created in titin serves to limit the range of motion of the sarcomere, giving rise to what is called passive stiffness.
Skeletal and cardiac muscles have slight amino acid sequence variations in their ti tin proteins and these appear to relate to differences in the mechanical characteristics of each muscle.
Energy backup for muscle energy
Myoglobin was described as a molecular batter for oxygen. Muscle cells have a better of their own for ATP. The is important for animals, but not for plants because a plant’s need for energy is different than an animal’s. Plants do not need to access energy sources as rapidly as animals do, nor do they have to maintain a constant internal temperature. Plants can neither flee predators, nor chase prey. These needs of animals are much more immediate and require that energy stores be accessible on demand. Muscles, of course, enable the motion of animals and the energy required for muscle contraction is ATP. To have stores of energy readily available, muscles have, in addition to ATP, creatine phosphate for energy and glycogen for quick release of glucose to make more energy. The synthesis of creatine phosphate is a prime example of the effects of concentration on the synthesis of high energy molecules. For example, creatine phosphate has an energy of hydrolysis of -43.1 kJ/mol whereas ATP has an energy of hydrolysis of -30.5 kJ/mol. Creatine phosphate, however, is made from creatine and ATP in the reaction shown in Figure 2.126. How is this possible?
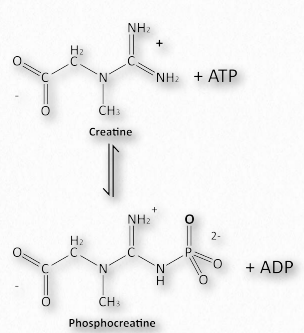
Figure 2.126 – Phosphorylation of creatine (phosphocreatine) – making of a creatine phosphate battery Image by Aleia Kim
The ∆G°’ of this reaction is +12.6 kJ/mol, reflecting the energies noted above. In a resting muscle cell, ATP is abundant and ADP is low, driving the reaction downward, creating creatine phosphate. When muscular contraction commences, ATP levels fall and ADP levels climb. The above reaction then reverses and proceeds to synthesize ATP immediately. Thus, creatine phosphate acts like a battery, storing energy when ATP levels are high and releasing it almost instantaneously to create ATP when its levels fall.

