104 Bacterial Transformation Lab
Genetically Modified Organisms
Dictionary.com defines a genetically modified organism (or GMO) as any organism whose genetic material has been altered using genetic engineering so that its DNA contains one or more genes not normally found there. GMOs are created by manipulating the genes of an organism to cause a change in the organism’s traits. GMOs are currently a hotly debated topic. Opponents of the technology claim that GMOs pose a health risk to humans as well as potential environmental risks. Supporters of GMOs believe that the benefits such as increasing the available food supply, increasing the nutritional content of foods, and the production of medicine outweigh the risks.
The glowing mice seen in Figure 1 are one example of a GMO. These mice contain the gene for green fluorescent protein, which was originally isolated from jellyfish. During this lab, you will be creating your own genetically modified organism by adding the green fluorescent protein gene to E. coli bacteria.

Bacterial Transformation
Transformation is the genetic alteration of a cell by the update of DNA from the environment. This process can occur naturally in some types of bacteria, but is typically rare. In a lab, we can subject bacteria to conditions that will cause them to take up DNA from the environment (to become “transformed”). There are several ways to transform bacteria in a lab setting, but one of the most common involves changing the concentration of ions in the bacteria’s surroundings and then heating the cells in a specific way. Bacteria that are able to easily take up DNA from the environment are called “competent”. Making cells competent renders their cell membrane more permeable to DNA. After the new DNA has entered the bacteria, it is used by the cell to make RNA and then protein. The new proteins produced from this DNA are what cause the change in the traits of the cells.
Plasmids
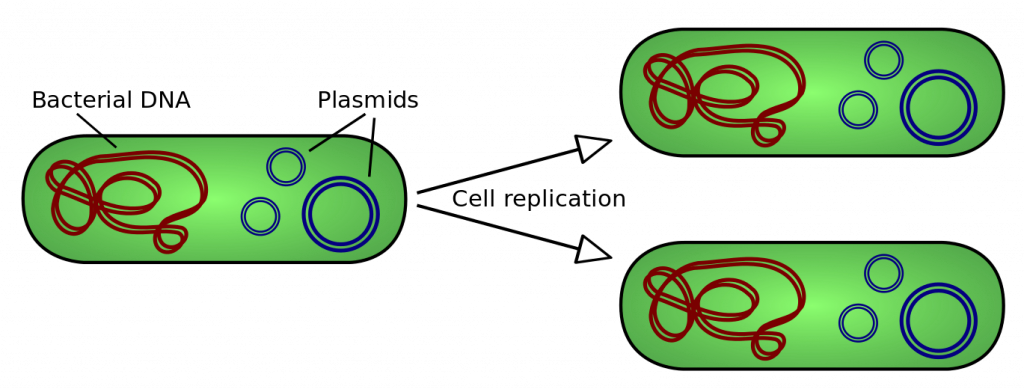
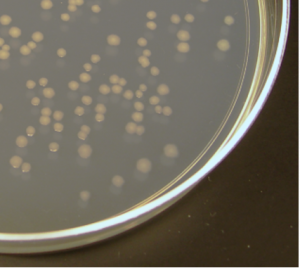
In addition to their DNA genome (which is circular), bacteria can also contain additional smaller circles of DNA called plasmids. A plasmid is a small, circular piece of double-stranded DNA that can be copied by bacterial cells. Plasmids occur naturally in bacteria and they are widely used by scientists as a method of for introducing foreign DNA into these cells because the sequence of DNA within the plasmid can be modified in the lab. Once a plasmid has entered a cell, it is copied by the cell’s DNA replication machinery. When the bacterial cell divides, each new daughter cell receives copies of the plasmid. One original transformed bacteria will divide to form a visible colony made up of one million or more transformed bacteria, which each contain a copy of the plasmid (Figure 3).
Selecting for transformed bacteria
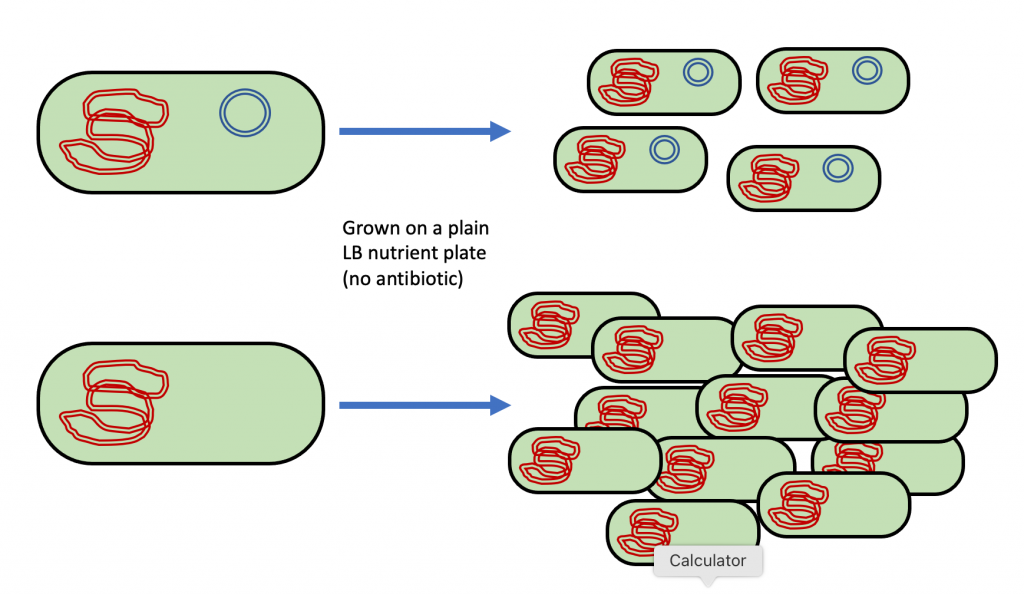
In order to transform bacteria using plasmid DNA, biotechnologists must overcome two problems. First, cells that contain plasmid DNA have a disadvantage since cellular resources (such as energy) are being used to replicate the plasmid and to synthesize the proteins that are encoded for by the plasmid’s DNA. If a mixed population of cells with plasmids and cells without plasmids is grown together in the presence of plenty of nutrients, then the cells without the plasmids grow faster because they are not wasting energy on a plasmid that they do not need (Figure 4). Therefore, there is always tremendous pressure on cells to get rid of their plasmids. If they are able to get rid of the plasmid, they will grow faster on a nutrient plate (or in the environment). However, getting rid of the plasmid is exactly what we do not want them to do. To overcome the pressure to get rid of the plasmid, we must provide an advantage to the cells that have and keep the plasmid.
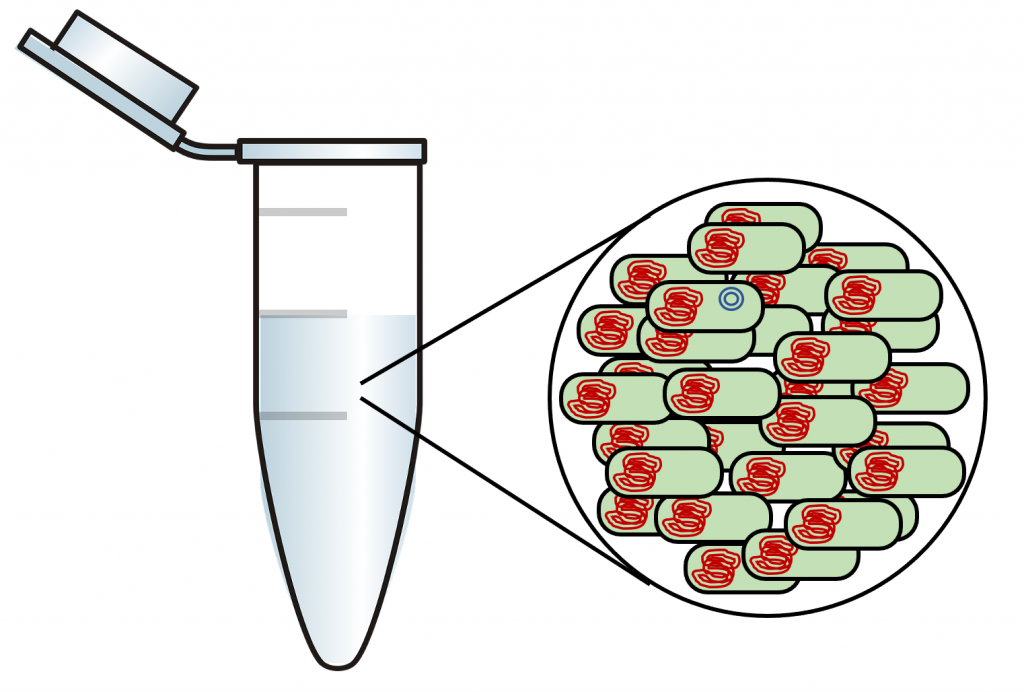
Second, we need to be able to determine which bacteria received the plasmid. In a typical transformation, billions of bacteria are treated to make them competent and then exposed to plasmid DNA. Typically, fewer than 1 in 1000 bacteria will acquire the plasmid (Figure 5). We need a way to get rid of the untransformed bacteria (greater than 99% of the total bacteria present) so that we are left with only the bacteria that were transformed with the plasmid. If we do not get rid of the untransformed bacteria, we will not be able to see the transformed bacteria since they are such a small percentage of the total number.
Antibiotic resistance genes provide a means of finding the bacteria that acquired the plasmid DNA in the midst of all those bacteria that did not. Antibiotics are chemicals that inhibit the growth of or kill bacteria. If the plasmid contains a gene for resistance to an antibiotic, then after transformation, bacteria grown on a nutrient plate containing the antibiotic will not be inhibited or killed by it. This means that bacteria that took up the plasmid during transformation can be distinguished from bacteria that did not by growing the bacteria on a nutrient plate containing the antibiotic (Figure 6). Only the bacteria that were transformed with the plasmid will survive the killing effect of the antibiotic and grow to form visible colonies on the plate. Remember that a colony is formed from more than one million genetically identical bacterial cells. This means that the only colonies growing on a nutrient + antibiotic plate after a transformation are the bacteria that acquired and kept the plasmid. Using an antibiotic in the nutrient plate and an antibiotic resistance gene in the plasmid accomplishes our two goals of giving an advantage to cells that have a plasmid so the plasmid is retained and of having a marker so we know our cells contain new DNA. Resistance to an antibiotic is known as a selectable marker because we can select for cells that contain it.
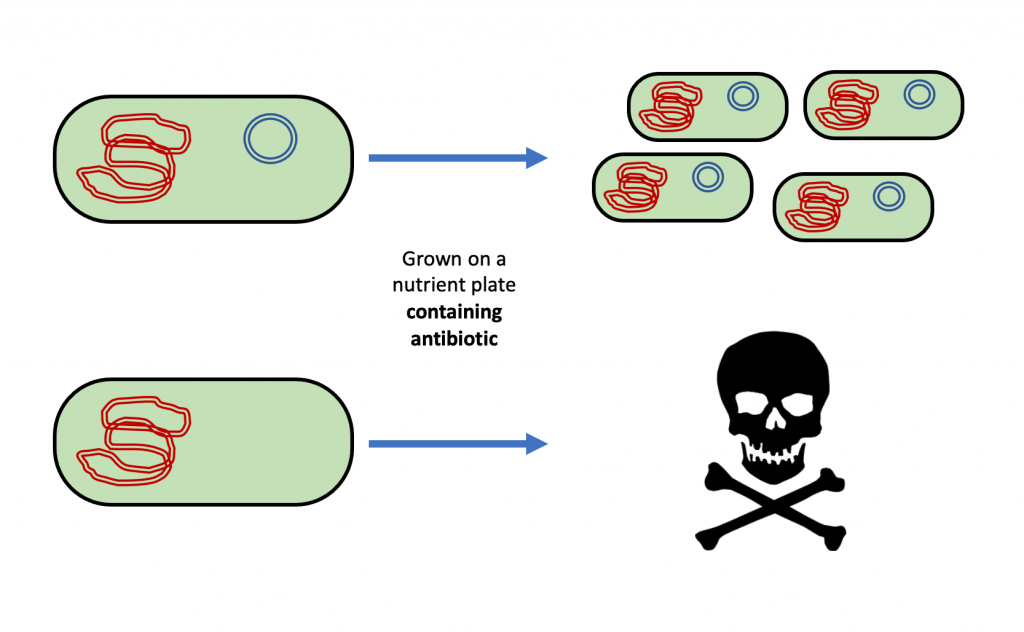
This is what it using an antibiotic to select transformed cells that contain a plasmid would look like:
Aseptic Technique
Aseptic technique is a set of methods that are used to prevent contamination. In a healthcare setting, it is used to prevent spreading dangerous microorganisms between patients. We will use aspects of aseptic technique during our bacterial transformation labs to prevent our plates and bacterial samples from becoming contaminated.
In a healthcare setting, you might wear gloves to prevent microorganisms from spreading. In this lab, it is not necessary to wear gloves since the bacteria that we are using are not all that dangerous, but you should wash your hands before you start the lab and again before you leave the lab room. You should also clean the surface of your lab bench at the start and end of lab, as directed by your instructor.
It is important that the sterile equipment and petri plates do not become contaminated due to contact with you or the environment. This means you should always open sterile equipment carefully, never leave it sitting open on your lab bench, and never set it down on a surface that is not sterile (for example, on the lab bench or on your lab notebook). This includes the lids of petri dishes or culture tubes – you should hold the lid above the petri dish over the surface of the dish rather than putting it on the lab bench. You should never touch the sterile portion of a piece of equipment with your hands before you use it, even if your hands are clean. For example, do not use your hands to put a sterile pipette tip on your pipettor, and only touch the handle end of an inoculating loop (not the loop end which will touch the bacteria).
Once a piece of equipment has touched bacteria, it is contaminated and must be discarded in the biohazard bag, not the regular lab trash can. You should collect all contaminated items (anything that has touched bacteria) in a waste beaker at your desk and discard them in the biohazard bag at the end of the lab. You should also never touch the something with your hands that has already touched bacteria (for example, pulling a used pipette tip off with your fingers).
NEVER PUT FOOD OR DRINK ON THE LAB BENCHES! If you need to access your water bottle (or coffee) during lab, wash your hands before you touch it.
For Our Lab:
The plasmid that we will be using is called pGLO (available from Bio-Rad). This plasmid contains several important pieces:
- Ori – an origin of replication, which allows the plasmid to be copied when the bacteria divide.
- GFP (green fluorescent protein) gene – the GFP protein gives a green glow in the presence of UV light.
- bla gene – the enzyme beta-lactamase is produced from this gene. This enzyme breaks down some antibiotics such as ampicillin when they are present in the environment before they can kill the bacteria.
- araC gene – the AraC protein produced by this gene turns on the GFP gene when arabinose is present in the environment.
Bacteria that are transformed with this plasmid will have two new traits: they will fluoresce green under UV light and they will be resistant to the antibiotic ampicillin.
The basic steps in the process of bacterial transformation are:
- Mix actively growing bacteria with the plasmid DNA you want to insert in a tube containing CaCl2 (calcium chloride) solution.
- “Heat shock” the bacteria by rapidly heating and then cooling them. This process causes the plasmid to enter the bacteria.
- Transfer the bacteria to an LB nutrient plate (containing nutrients) so that they can recover and express their newly acquired genes.
After the bacterial transformation procedure has been carried out, cells that contain the plasmid are selected for by growing the bacteria on LB nutrient plates that contain ampicillin. The ampicillin kills any cell that did not get transformed with the plasmid. This means that the only bacteria which can grow to form visible colonies on a plate containing LB nutrients and ampicillin are transformed cells. These cells will produce GFP at very low levels and will appear whitish when viewed under UV light.
Arabinose is a type of sugar that can be added to the plates when they are poured. Although arabinose is a sugar, it is not being used as a nutrient source in this experiment. When transformed bacteria are grown on plates containing LB nutrients + ampicillin + arabinose, the arabinose interacts with the araC protein (which is produced from the araC gene). The interaction of arabinose + araC protein stimulates transcription of the GFP gene. This results in a brilliant green glow when the bacteria are viewed under a UV light source.
Samples in our experiment
In our lab, we will compare transformed (+pGLO) and non-transformed (-pGLO) bacteria grown on several different types of plate. Here are the key points to remember:
- Ampicillin kills bacteria that do not contain the bla gene. This means that only bacteria that contain the pGLO plasmid (on which the bla gene is found) will grow on plates containing the antibiotic ampicillin.
- The GFP gene is located on the pGLO plasmid. This means that only bacteria that contain the pGLO plasmid can fluoresce green under UV light.
- Arabinose turns on the expression of the GFP gene. This means that bacteria that contain the pGLO plasmid will only fluoresce in the presence of arabinose.
Determine which plate below matches with the label to the right.
Identify which of these statements are true about each of the materials used in this experiment by moving the YES to the appropriate boxes.

