47 Gene Regulation
Each cell expresses, or turns on, only a fraction of its genes. “Expresses” or “turns on” means that protein is being produced from that gene. The rest of the genes are repressed, or turned off (no protein is being produced from those genes). The process of turning genes on and off is known as gene regulation. Gene regulation is an important part of normal development. Genes are turned on and off in different patterns during development to make a brain cell look and act different from a liver cell or a muscle cell, for example. Gene regulation also allows cells to react quickly to changes in their environments. Although we know that the regulation of genes is critical for life, this complex process is not yet fully understood.
For a cell to function properly, necessary proteins must be synthesized at the proper time. All organisms and cells control or regulate the transcription and translation of their DNA into protein. The process of turning on a gene to produce RNA and protein is called gene expression. Whether in a simple unicellular organism or in a complex multicellular organism, each cell controls when and how its genes are expressed. For this to occur, there must be a mechanism to control when a gene is expressed to make RNA and protein, how much of the protein is made, and when it is time to stop making that protein because it is no longer needed.
Cells in multicellular organisms are specialized; cells in different tissues look very different and perform different functions. For example, a muscle cell is very different from a liver cell, which is very different from a skin cell. These differences are a consequence of the expression of different sets of genes in each of these cells. All cells have certain basic functions they must perform for themselves, such as converting the energy in sugar molecules into energy in ATP. Therefore, there is a set of “housekeeping” genes that are expressed in all cells. Each type of cell also has many genes that are not expressed because the cell does not need to perform those functions. Specific cells also express many genes that are not expressed by other cells so that they can carry out their specialized functions. In addition, cells will turn on or off certain genes at different times in response to changes in the environment or at different times during the development of the organism. Unicellular organisms, both eukaryotic and prokaryotic, also turn on and off genes in response to the demands of their environment so that they can respond to special conditions.

The control of gene expression is extremely complex. Malfunctions in this process are detrimental to the cell and can lead to the development of many diseases, including cancer.
Epigenetic Regulation
DNA modifications that do not change the DNA sequence can affect gene activity. Chemical compounds that are added to single genes can regulate their activity; these modifications are known as epigenetic changes. The epigenome comprises all of the chemical compounds that have been added to the entirety of one’s DNA (genome) as a way to regulate the activity (expression) of all the genes within the genome. The chemical compounds of the epigenome are not part of the DNA sequence, but are on or attached to DNA (“epi-“ means above in Greek). Epigenomic modifications remain as cells divide and in some cases can be inherited through the generations. Environmental influences, such as a person’s diet and exposure to pollutants, can also impact the epigenome.
Epigenetic changes can help determine whether genes are turned on or off and can influence the production of proteins in certain cells, ensuring that only necessary proteins are produced. For example, proteins that promote bone growth are not produced in muscle cells. Patterns of epigenome modification vary among individuals, different tissues within an individual, and even different cells.
The human genome encodes over 20,000 genes, which means that each of the 23 pairs of human chromosomes contains thousands of genes. The DNA in the nucleus of each cell is precisely wound, folded, and compacted into chromosomes so that it will fit inside the nuclear membrane. It is also organized so that specific segments can be accessed as needed by a specific cell type.
The first level of organization, or packing, is the winding of DNA strands around histone proteins. Histones package and order DNA into structural units called nucleosome complexes, which can control the access of proteins to the DNA regions (Figure 1a). Under the electron microscope, this winding of DNA around histone proteins to form nucleosomes looks like small beads on a string (Figure 1b). These beads (histone proteins) can move along the string (DNA) and change the structure of the molecule.
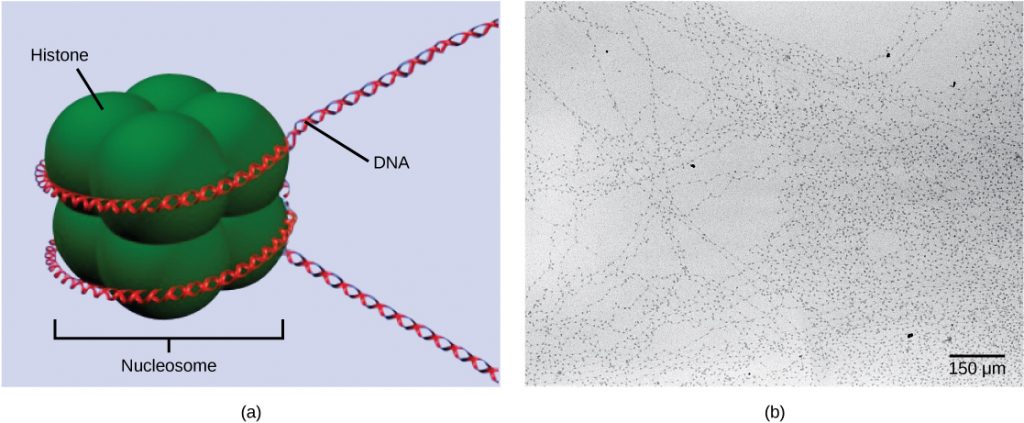
If DNA encoding a specific gene is to be transcribed into RNA, the nucleosomes surrounding that region of DNA can slide down the DNA to open that specific chromosomal region and allow for the transcriptional machinery (RNA polymerase) to initiate transcription. Nucleosomes can move to open the chromosome structure to expose a segment of DNA, but do so in a very controlled manner.
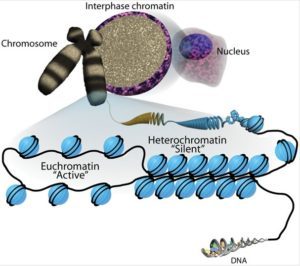
Active open regions of chromatin are called euchromatin (Figure 2). Regions of the genome that are transcriptionally active are typically euchromatic. Tightly wound regions of chromatin are called heterochromatin. Heterochromatic regions of the genome are typically silenced and transcriptionally inactive.
Modifications to DNA and histones
How the histone proteins move, and whether the DNA is wrapped loosely or tightly around them, is dependent on signals found on both the histone proteins and on the DNA. These signals are chemical tags added to histone proteins and DNA that tell the histones if a chromosomal region should be open or closed. These tags are not permanent, but may be added or removed as needed. They are chemical modifications (phosphate, methyl, or acetyl groups) that are attached to specific amino acids in the protein or to the nucleotides of the DNA. The tags do not alter the DNA base sequence, but they do alter how tightly wound the DNA is around the histone proteins.
This type of gene regulation is called epigenetic regulation. Epigenetic means “around or above genetics.” The changes that occur to the histone proteins and DNA do not alter the nucleotide sequence and are not permanent. Instead, these changes are temporary, although they can and often do persist through multiple rounds of cell division. They alter the chromosomal structure (open euchromatin or closed heterochromatin) as needed, but do not change the sequence of bases within the DNA.
A gene can be turned on or off depending upon the location and modifications to the histone proteins and DNA. If a gene is to be transcribed, the histone proteins and DNA are modified surrounding the chromosomal region encoding that gene. This opens the chromosomal region (it becomes euchromatic) to allow access for RNA polymerase and other proteins, called transcription factors, to bind to the promoter region, located just upstream of the gene, and initiate transcription. If a gene is to remain turned off, or silenced, the histone proteins and DNA have different modifications that signal a closed chromosomal configuration. In this closed configuration (heterochromatin), the RNA polymerase and transcription factors do not have access to the DNA and transcription cannot occur (Figure 2).
DNA Methylation
A common type of epigenomic modification is called methylation.Methylation involves attaching small molecules called methyl groups, each consisting of one carbon atom and three hydrogen atoms, to DNA nucleotides or the amino acids that make up the histone proteins.
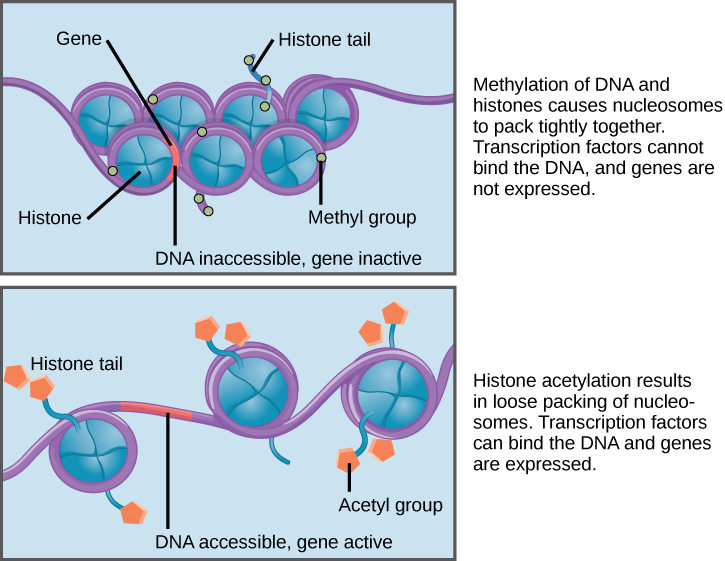
When DNA is methylated, the methyl group is typically added to cytosine nucleotides. This occurs within very specific regions called CpG islands. These are stretches with a high frequency of cytosine and guanine dinucleotide DNA pairs (CG) found in the promoter regions of genes. When this configuration exists, the cytosine member of the pair can be methylated (a methyl group is added). This modification changes how the DNA interacts with proteins, including the histone proteins that control access to the region. When methyl groups are added to a particular gene, that gene is turned off or silenced, and no protein is produced from that gene (Figure 3).
“Histone Code” Hypothesis
The histone code hypothesis is the hypothesis that transcription of a gene is in part regulated by modifications made to histone proteins, primarily on their somewhat floppy ends (their “tails”). Many of the histone tail modifications correlate very well to chromatin structure and both histone modification state and chromatin structure correlate well to gene expression levels. The most important concept in the histone code hypothesis is that the histone modifications serve to recruit other proteins by specific recognition of the modified histone, rather than through simply stabilizing or destabilizing the interaction between histone and the underlying DNA. These recruited proteins then act to alter chromatin structure actively or to promote transcription.
The histone code has the potential to be massively complex. There are at least 20 modifications that are made to histone tails that have been relatively well characterized, and there is the potential for many more that we have not discovered. Each histone can be modified on multiple amino acids, with multiple different chemical modifications. The information that can be stored in the histone code dwarfs the amount that is stored in the order of the bases in the human genome.
Histone Methlyation
A portion of the histone protein known as the histone tail can have methyl groups (CH3) added to it. This is the same modification that is made to cytosine nucleotides in DNA. The specific amino acid in the histone tail that gets methylated is very important for determining whether it will tighten or loosen chromatin structure. Modification to several amino acids in the tail is correlated with euchromatin and active transcription, while modification to other amino acids is correlated with heterochromatin and gene silencing. You should know that histones can be methylated, but we can’t use histone methylation as a predictor for euchromatin or heterochromatin.
Histone Acetylation
Histone tails can also be modified by the addition of an acetyl group (this process is known as acetylation). If you remember from cellular respiration, an acetyl group (such as that found in acetyl-CoA) is a 2-carbon molecule. When histone tails are acetylated, this typically causes the tails to loosen from around the DNA, allowing the chromatin to loosen (Figure 3).
Other modifications
There are many other modifications that can be made to histone proteins in addition to methylation and acetylation. Histone tails can be phosphorylated or ubiquitinated (where a small protein called ubiquitin is attached). Histone phosphorylation seems to be related to DNA repair. Ubiquitination has been shown to be associated with both transcriptional activation or inactivation, depending on the specific location.
Epigenetic Changes
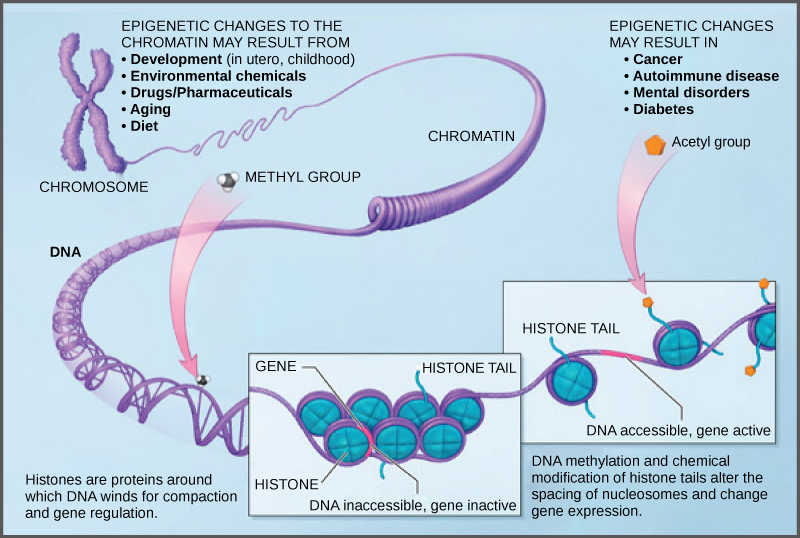
Because errors in the epigenetic process, such as modifying the wrong gene or failing to add a compound to a gene, can lead to abnormal gene activity or inactivity, they can cause genetic disorders. Conditions including cancers, metabolic disorders, and degenerative disorders have all been found to be related to epigenetic errors.
Cancerous cells often have regions of DNA that show different levels of methylation compared to normal cells. Some genes are methylated and silenced in cancerous cells, while they are unmethylated and active in normal cells. Other genes are active in cancerous cells, but inactive in normal cells. Each specific cancer in each specific individual can show different patterns of methylation, although there are similarities between many different types of cancer.
Scientists continue to explore the relationship between the genome and the chemical compounds that modify it. In particular, they are studying what effect the modifications have on gene function, protein production, and human health.
Transcriptional Regulation
Gene expression can be regulated at the transcriptional level. This means that the process of transcription can be turned on or off. In both prokaryotic and eukaryotic cells, transcription requires RNA polymerase to bind to a sequence upstream of a gene to initiate transcription. Prokaryotes almost always regulate gene expression at the transcriptional level.
In eukaryotes, the eukaryotic RNA polymerase requires other proteins, or transcription factors, to facilitate transcription initiation. Transcription factors are proteins that bind to the promoter sequence and other regulatory sequences to control the transcription of the target gene. RNA polymerase by itself cannot initiate transcription in eukaryotic cells. Transcription factors must bind to the promoter region first and recruit RNA polymerase to the site for transcription to be established.
- If transcription factors are not allowed to bind, transcription can not take place, which means gene expression is turned off.
- In some eukaryotic genes, there are regions that help increase or enhance transcription called enhancers. When transcription factors bind to these enhancer regions, they can increase rates of transcription, which means gene expression is turned up.
- Transcriptional repressors can bind to promoter or enhancer regions and block transcription. This means that gene expression is turned off.
Post-transcriptional Regulation
After RNA is transcribed, it must be processed into a mature form before translation can begin. This processing after an RNA molecule has been transcribed, but before it is translated into a protein, is called post-transcriptional modification. As with the epigenetic and transcriptional stages of processing, this post-transcriptional step can also be regulated to control gene expression in the cell. If the RNA is not processed, shuttled, or translated, then no protein will be synthesized.
Alternative RNA Splicing
In the 1970s, genes were first observed that exhibited alternative RNA splicing. Alternative RNA splicing is a mechanism that allows different protein products to be produced from one gene when different combinations of introns (and sometimes exons) are removed from the transcript (Figure 1). This alternative splicing can be haphazard, but more often it is controlled and acts as a mechanism of gene regulation, with the frequency of different splicing alternatives controlled by the cell as a way to control the production of different protein products in different cells, or at different stages of development. Alternative splicing is now understood to be a common mechanism of gene regulation in eukaryotes; according to one estimate, 70% of genes in humans are expressed as multiple proteins through alternative splicing.
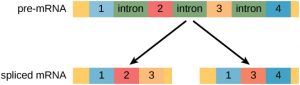
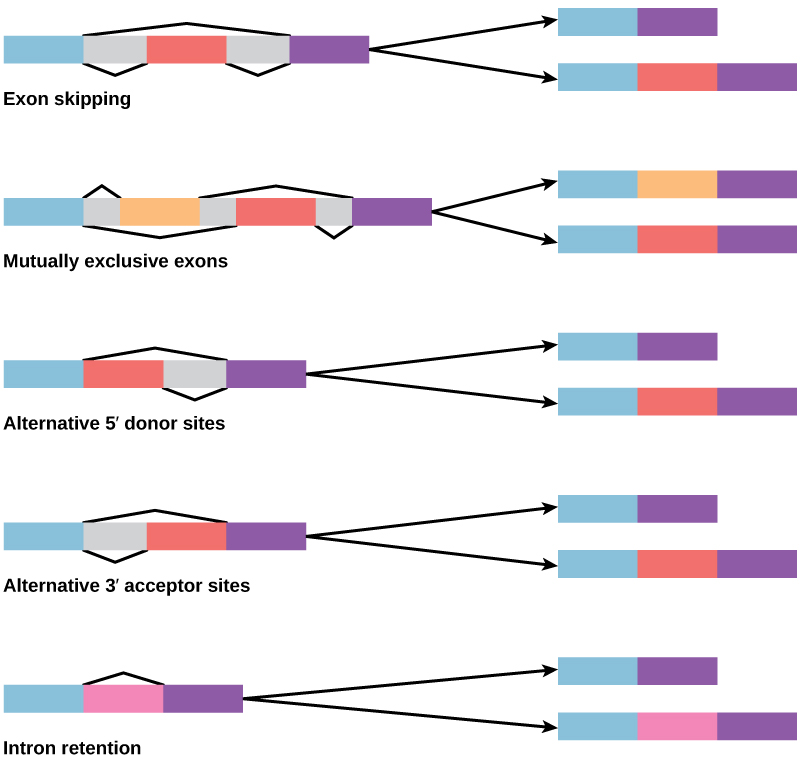
How could alternative splicing evolve? Introns have a beginning and ending recognition sequence, and it is easy to imagine the failure of the splicing mechanism to identify the end of an intron and find the end of the next intron, thus removing two introns and the intervening exon. In fact, there are mechanisms in place to prevent such exon skipping, but mutations are likely to lead to their failure. Such “mistakes” would more than likely produce a nonfunctional protein. Indeed, the cause of many genetic diseases is alternative splicing rather than mutations in a sequence. However, alternative splicing would create a protein variant without the loss of the original protein, opening up possibilities for adaptation of the new variant to new functions. Gene duplication has played an important role in the evolution of new functions in a similar way—by providing genes that may evolve without eliminating the original functional protein.
Control of RNA Stability
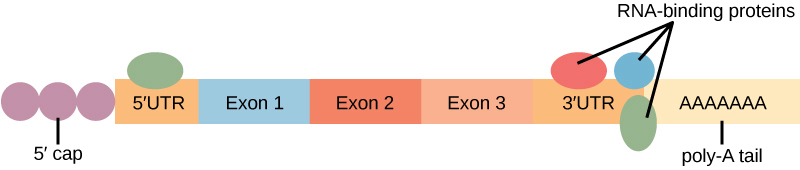
Before the mRNA leaves the nucleus, it is given two protective “caps” that prevent the end of the strand from degrading during its journey. The 5′ cap, which is placed on the 5′ end of the mRNA, is usually composed of a methylated guanosine triphosphate molecule (GTP). The poly-A tail, which is attached to the 3′ end, is usually composed of a series of adenine nucleotides. Once the RNA is transported to the cytoplasm, the length of time that the RNA remains there can be controlled. Each RNA molecule has a defined lifespan and decays at a specific rate. This rate of decay can influence how much protein is in the cell. If the RNA decays more rapidly, translation has less time to occur, so less protein will be produced. Conversely, if RNA decays less rapidly, more protein will be produced. This rate of decay is referred to as the RNA stability. If the RNA is stable, it will be detected for longer periods of time in the cytoplasm. Binding of proteins to the RNA can influence its stability (Figure 3).
Translational Regulation
Like transcription, translation is controlled by proteins that bind and initiate the process. In translation, the complex that assembles to start the process is referred to as the initiation complex. Regulation of the formation of this complex can increase or decrease rates of translation.
Post-translational Regulation
Proteins can be chemically modified with the addition of groups including methyl, phosphate, acetyl, and ubiquitin groups. The addition or removal of these groups from proteins regulates their activity or the length of time they exist in the cell. Sometimes these modifications can regulate where a protein is found in the cell—for example, in the nucleus, the cytoplasm, or attached to the plasma membrane.
Chemical modifications occur in response to external stimuli such as stress, the lack of nutrients, heat, or ultraviolet light exposure. These changes can alter epigenetic accessibility, transcription, mRNA stability, or translation—all resulting in changes in expression of various genes. This is an efficient way for the cell to rapidly change the levels of specific proteins in response to the environment. Because proteins are involved in every stage of gene regulation, the phosphorylation of a protein (depending on the protein that is modified) can alter accessibility to the chromosome, can alter translation (by altering transcription factor binding or function), can change nuclear shuttling (by influencing modifications to the nuclear pore complex), can alter RNA stability (by binding or not binding to the RNA to regulate its stability), can modify translation (increase or decrease), or can change post-translational modifications (add or remove phosphates or other chemical modifications).
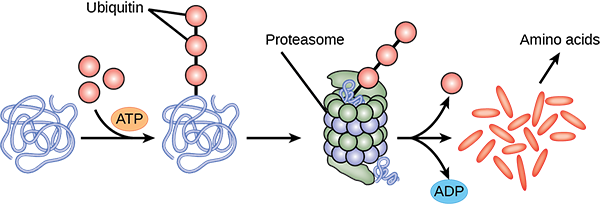
The addition of an ubiquitin group to a protein marks that protein for degradation. Ubiquitin acts like a flag indicating that the protein lifespan is complete. These proteins are moved to the proteasome, an organelle that functions to remove proteins, to be degraded (Figure 2). One way to control gene expression, therefore, is to alter the longevity of the protein.
References
Unless otherwise noted, images on this page are licensed under CC-BY 4.0 by OpenStax.
OpenStax, Concepts of Biology. OpenStax CNX. January 3, 2017. https://cnx.org/contents/GFy_h8cu@10.120:7Ry3oRse@6

