Eukaryotic Transcription
Prokaryotes and eukaryotes perform fundamentally the same process of transcription, with a few key differences. The most important difference between prokaryotes and eukaryotes is the latter’s membrane-bound nucleus and organelles. With the genes enclosed in a nucleus, the eukaryotic cell must be able to transport its mRNA to the cytoplasm and must protect its mRNA from degrading before it is translated. Eukaryotes also employ three different polymerases that each transcribe a different subset of genes.
Initiation
The eukaryotic promoters that we are most interested in are similar to prokaryotic promoters in that they contain a TATA box (Figure 1). However, initiation of transcription is much more complex in eukaryotes compared to prokaryotes. Unlike the prokaryotic RNA polymerase that can bind to a DNA template on its own, eukaryotes require several other proteins, called transcription factors, to first bind to the promoter region and then help recruit the appropriate polymerase.
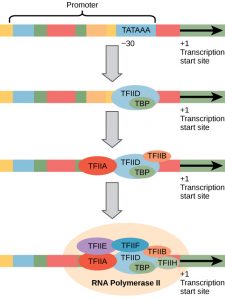
In addition, there are three different RNA polymerases in eukaryotes, each of which is made up of 10 subunits or more. Each eukaryotic RNA polymerase also requires a distinct set of transcription factors to bring it to the DNA template.
RNA polymerase I is located in the nucleolus, a specialized nuclear substructure in which ribosomal RNA (rRNA) is transcribed, processed, and assembled into ribosomes. The rRNA molecules are considered structural RNAs because they have a cellular role but are not translated into protein. The rRNAs are components of the ribosome and are essential to the process of translation. RNA polymerase I synthesizes most of the rRNAs.
RNA polymerase II is located in the nucleus and synthesizes all protein-coding nuclear pre-mRNAs. Eukaryotic pre-mRNAs undergo extensive processing after transcription but before translation. For clarity, the term “mRNA” will only be used to describe the mature, processed molecules that are ready to be translated. RNA polymerase II is responsible for transcribing the overwhelming majority of eukaryotic genes.
RNA polymerase III is also located in the nucleus. This polymerase transcribes a variety of structural RNAs including transfer pre-RNAs (pre-tRNAs), and small nuclear pre-RNAs. The tRNAs have a critical role in translation; they serve as the adaptor molecules between the mRNA template and the growing polypeptide chain. Small nuclear RNAs have a variety of functions, including “splicing” pre-mRNAs and regulating transcription factors.
Each of the types of RNA polymerase recognizes a different promoter sequence and requires different transcription factors.
Elongation
Following the formation of the preinitiation complex, the polymerase is released from the other transcription factors, and elongation is allowed to proceed as it does in prokaryotes with the RNA polymerase synthesizing pre-mRNA in the 5′ to 3′ direction. As discussed previously, RNA polymerase II transcribes the major share of eukaryotic genes, so this section will focus on how this polymerase accomplishes elongation and termination.
Although the enzymatic process of elongation is essentially the same in eukaryotes and prokaryotes, the DNA template is more complex. When eukaryotic cells are not dividing, their genes exist as a diffuse mass of DNA and proteins called chromatin. The DNA is tightly packaged around charged histone proteins at repeated intervals. These DNA–histone complexes, collectively called nucleosomes, are regularly spaced and include 146 nucleotides of DNA wound around eight histones like thread around a spool.
For RNA synthesis to occur, the transcription machinery needs to move histones out of the way every time it encounters a nucleosome. This is accomplished by a special protein complex called FACT, which stands for “facilitates chromatin transcription.” This complex pulls histones away from the DNA template as the polymerase moves along it. Once the pre-mRNA is synthesized, the FACT complex replaces the histones to recreate the nucleosomes.
Termination
The termination of transcription is different for the different polymerases. Unlike in prokaryotes, elongation by RNA polymerase II in eukaryotes takes place 1,000–2,000 nucleotides beyond the end of the gene being transcribed. This pre-mRNA tail is removed during mRNA processing. RNA polymerases I and III require termination signals. Genes transcribed by RNA polymerase I contain a specific 18-nucleotide sequence that is recognized by a termination protein. The process of termination in RNA polymerase III involves an mRNA hairpin that causes the mRNA to be released.
References
Unless otherwise noted, images on this page are licensed under CC-BY 4.0 by OpenStax.
OpenStax, Biology. OpenStax CNX. December 21, 2017. https://cnx.org/contents/GFy_h8cu@10.120:6l70P9u6@5/Eukaryotic-Transcription

